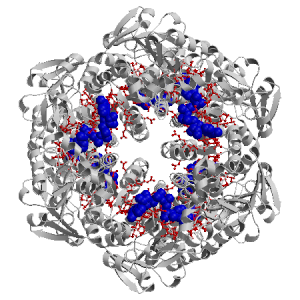
| NAD B 500 |
|---|
| 129 | A |
LEU |
| 132 | A |
MET |
| 133 | A |
SER |
| 136 | A |
ALA |
| 173 | A |
LEU |
| 174 | A |
GLY |
| 175 | A |
GLY |
| 176 | A |
GLY |
| 177 | A |
VAL |
| 178 | A |
VAL |
| 196 | A |
PHE |
| 197 | A |
ASP |
| 198 | A |
ILE |
| 199 | A |
ASN |
| | 202 | A |
ARG |
| 219 | A |
SER |
| 237 | A |
ALA |
| 238 | A |
VAL |
| 239 | A |
LEU |
| 240 | A |
VAL |
| 246 | A |
PRO |
| 248 | A |
LEU |
| 266 | A |
VAL |
| 267 | A |
ALA |
| 268 | A |
VAL |
| 269 | A |
ASP |
| 270 | A |
GLN |
| 298 | A |
PRO |
| 299 | A |
ASN |
| | 300 | A |
MET |
| 301 | A |
PRO |
| NAD H 500 |
|---|
| 129 | G |
LEU |
| 132 | G |
MET |
| 133 | G |
SER |
| 136 | G |
ALA |
| 173 | G |
LEU |
| 174 | G |
GLY |
| 175 | G |
GLY |
| 176 | G |
GLY |
| 177 | G |
VAL |
| 178 | G |
VAL |
| 196 | G |
PHE |
| 197 | G |
ASP |
| | 198 | G |
ILE |
| 199 | G |
ASN |
| 202 | G |
ARG |
| 219 | G |
SER |
| 237 | G |
ALA |
| 238 | G |
VAL |
| 239 | G |
LEU |
| 240 | G |
VAL |
| 246 | G |
PRO |
| 248 | G |
LEU |
| 266 | G |
VAL |
| 267 | G |
ALA |
| 268 | G |
VAL |
| 269 | G |
ASP |
| 270 | G |
GLN |
| | 298 | G |
PRO |
| 299 | G |
ASN |
| 300 | G |
MET |
| 301 | G |
PRO |
| NAD F 500 |
|---|
| 132 | E |
MET |
| 133 | E |
SER |
| 136 | E |
ALA |
| 173 | E |
LEU |
| 174 | E |
GLY |
| 175 | E |
GLY |
| 176 | E |
GLY |
| 177 | E |
VAL |
| 178 | E |
VAL |
| 196 | E |
PHE |
| | 197 | E |
ASP |
| 198 | E |
ILE |
| 199 | E |
ASN |
| 202 | E |
ARG |
| 219 | E |
SER |
| 237 | E |
ALA |
| 238 | E |
VAL |
| 239 | E |
LEU |
| 240 | E |
VAL |
| 246 | E |
PRO |
| 248 | E |
LEU |
| 266 | E |
VAL |
| 267 | E |
ALA |
| 268 | E |
VAL |
| 269 | E |
ASP |
| | 270 | E |
GLN |
| 298 | E |
PRO |
| 299 | E |
ASN |
| 300 | E |
MET |
| 301 | E |
PRO |
| NAD D 500 |
|---|
| 132 | C |
MET |
| 133 | C |
SER |
| 136 | C |
ALA |
| 173 | C |
LEU |
| 174 | C |
GLY |
| 175 | C |
GLY |
| 176 | C |
GLY |
| 177 | C |
VAL |
| 178 | C |
VAL |
| | 196 | C |
PHE |
| 197 | C |
ASP |
| 198 | C |
ILE |
| 199 | C |
ASN |
| 202 | C |
ARG |
| 219 | C |
SER |
| 237 | C |
ALA |
| 238 | C |
VAL |
| 239 | C |
LEU |
| 240 | C |
VAL |
| 246 | C |
PRO |
| 248 | C |
LEU |
| 266 | C |
VAL |
| 267 | C |
ALA |
| 268 | C |
VAL |
| | 269 | C |
ASP |
| 270 | C |
GLN |
| 298 | C |
PRO |
| 299 | C |
ASN |
| 300 | C |
MET |
| 301 | C |
PRO |
| NAD J 500 |
|---|
| 132 | I |
MET |
| 133 | I |
SER |
| 136 | I |
ALA |
| 173 | I |
LEU |
| 174 | I |
GLY |
| 175 | I |
GLY |
| 176 | I |
GLY |
| 177 | I |
VAL |
| | 178 | I |
VAL |
| 196 | I |
PHE |
| 197 | I |
ASP |
| 198 | I |
ILE |
| 199 | I |
ASN |
| 202 | I |
ARG |
| 219 | I |
SER |
| 237 | I |
ALA |
| 238 | I |
VAL |
| 239 | I |
LEU |
| 240 | I |
VAL |
| 246 | I |
PRO |
| 248 | I |
LEU |
| 266 | I |
VAL |
| 267 | I |
ALA |
| | 268 | I |
VAL |
| 269 | I |
ASP |
| 270 | I |
GLN |
| 298 | I |
PRO |
| 299 | I |
ASN |
| 300 | I |
MET |
| 301 | I |
PRO |
| NAD L 500 |
|---|
| 132 | K |
MET |
| 133 | K |
SER |
| 136 | K |
ALA |
| 173 | K |
LEU |
| 174 | K |
GLY |
| 175 | K |
GLY |
| 176 | K |
GLY |
| | 177 | K |
VAL |
| 178 | K |
VAL |
| 196 | K |
PHE |
| 197 | K |
ASP |
| 198 | K |
ILE |
| 199 | K |
ASN |
| 202 | K |
ARG |
| 219 | K |
SER |
| 237 | K |
ALA |
| 238 | K |
VAL |
| 239 | K |
LEU |
| 240 | K |
VAL |
| 246 | K |
PRO |
| 248 | K |
LEU |
| 266 | K |
VAL |
| | 267 | K |
ALA |
| 268 | K |
VAL |
| 269 | K |
ASP |
| 270 | K |
GLN |
| 298 | K |
PRO |
| 299 | K |
ASN |
| 300 | K |
MET |
| 301 | K |
PRO |
|