 |
LigASite database of binding sites |
|
PDB ID and HEADER, TITLE and
COMPND records of the PDB file. | | (click anywhere in this window to remove it) |
|
| 1e65 |
| ELECTRON TRANSPORT(COPPER BINDING) |
HEADER |
|
|
| AZURIN FROM PSEUDOMONAS AERUGINOSA, APO FORM |
TITLE |
|
|
|
|
|
|
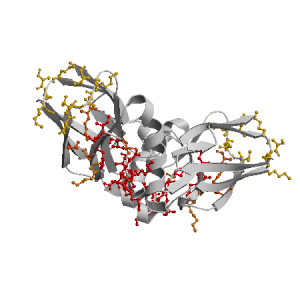
Figure highlighting the binding site residues. Figures were drawn with
Molscript (7) and rendered with
Raster3D (8). PISA coordinates
(3) are used when available
(all entries except NMR). | | (click anywhere in this window to remove it) |
|
 |
|
List of binding site residues detected in this protein. Column 1 gives the position, coloured on a yellow-to-red scale depending on the fraction of corresponding holo-structures where the residue is in contact with a ligand.
Column 2 gives the 3-letter amino acid code, coloured according to physico-chemical type. Chain ID's of residues are not mentioned in this page because all chains in the apo-structure refer to the same protein. | | (click anywhere in this window to remove it) |
|
| 24 | |
LYS |
| 25 | |
SER |
| 26 | |
CYS |
| 27 | |
LYS |
| 70 | |
LYS |
| 71 | |
ASP |
| 73 | |
LEU |
| 74 | |
LYS |
| 75 | |
PRO |
| 76 | |
ASP |
| 77 | |
ASP |
| 78 | |
SER |
| 80 | |
VAL |
| 81 | |
ILE |
| 82 | |
ALA |
| | 83 | |
HIS |
| 84 | |
THR |
| 85 | |
LYS |
| 95 | |
VAL |
| 98 | |
ASP |
| 99 | |
VAL |
| 100 | |
SER |
| 101 | |
LYS |
| 102 | |
LEU |
| 103 | |
LYS |
| 127 | |
LEU |
| 270 | |
LYS |
| 271 | |
ASP |
| 273 | |
LEU |
| 274 | |
LYS |
| | 275 | |
PRO |
| 276 | |
ASP |
| 277 | |
ASP |
| 280 | |
VAL |
| 281 | |
ILE |
| 282 | |
ALA |
| 283 | |
HIS |
| 295 | |
VAL |
| 301 | |
LYS |
|
|
|
| PDB |
The Protein Data Bank |
| CSA |
Catalytic Site Atlas |
| PDBSum |
Overview of the macromolecular structure |
| CATH |
Protein Structure Classification |
| Scop |
Structural Classification of Proteins |
| Pfam |
Protein Families and Domains |
| UniProt |
Universal Protein Resource |
LIGPLOT (only on holo-pages) is hosted at the EBI. The LigPlot Jmol links point directly to the Jmol visualisation interface provided on the PDBSum page. Note that due to different software used, the atomic contacts of LigPlot and LigASite do not necessarily correspond. | | (click anywhere in this window to remove it) |
|
Links to external databases: |
|
Several files are provided for download: | • The XML file defining the residue-ligand contacts; this file contains data on the apo and all holo-structures. |
| • The XML Schema file defining the semantics of the XML file |
| • 3D coordinates of the structure used in constructing LigASite (PISA structure file whenever available, PDB file otherwise. |
| • 3D coordinates of the combined binding residues in the apo structure |
| • 3D coordinates of the binding residues of the holo structure (only on the holo page) |
Coordinate files are in PDB format. | | (click anywhere in this window to remove it) |
|
|
|
|
|
Table describing the holo-structures and ligands used to define
the binding sites.
Column 1 gives the PDB ID of the holo-structure.
Column 2 gives the unique ID of the ligand;
a space-separated list of HET-groups that constitute
the ligand (see Methods).
Each HET-group in the ligand is uniquely identified by
a string in which the first four characters are the three-letter
HET ID from the PDB file followed by the chain ID from
the PISA file, and the last four characters are the residue sequence
number from the PDB file.
Column 3 gives the number of atoms in each ligand.
Column 4 gives the number of protein-ligand inter-atomic
contacts. | | (click anywhere in this window to remove it) |
|
| pdb ID |
Ligand Unique ID |
#atoms |
#contacts |
| 1jzg |
RTBB_903 |
33 |
76 |
Details |
|
RTBD_903 |
33 |
78 |
| 1jzf |
RTBB_903 |
33 |
77 |
Details |
|
RTBD_903 |
33 |
75 |
| 1jzh |
RTAA_902 |
31 |
73 |
Details |
| 1bex |
RBUD_130 |
30 |
82 |
Details |
|
RBUB_130 |
30 |
77 |
| 1jzj |
IMEJ2000 |
23 |
71 |
Details |
|
DOSN_905 |
30 |
72 |
|
IMEB2000 |
23 |
71 |
|
DOSF_905 |
30 |
72 |
|
DOSD_903 |
30 |
75 |
|
DOSL_903 |
30 |
75 |
|
|
|
|
v9.7
April 2012 |
Interdisciplinary Research Institute, Computational Biology, Villeneuve d'Ascq, France
University College London, Biomolecular Structure and Modelling Unit, London, UK
Hospital for Sick Children and University of Toronto, Structural Biology and Biochemistry Program, Toronto, Canada |
| Script execution time: 0.0681 seconds |

